ANY OpenSees from a Notebook#
Running OpenSees & OpenSeesPy Input Scripts from a DesignSafe Jupyter Notebook
by Silvia Mazzoni, DesignSafe, 2025
This notebook shows how to run any OpenSees script — whether written in Tcl or Python (OpenSeesPy) — using Python’s built-in os library to execute commands just like in a terminal.
You can apply this technique in:
A Jupyter notebook, for an interactive and user-friendly experience
A standalone Python script, for automated or large-scale workflows
Why Use This Approach?#
It makes running OpenSees in the DesignSafe JupyterHub environment highly user-friendly
It lets you build seamless, Python-driven workflows that can include preprocessing, simulation, and postprocessing in one place
It works with any version of OpenSees already available in the Jupyter environment
You can also apply the same method to call OpenSees on an HPC system (e.g., Stampede3), making it easier to scale up or integrate into continuous workflows
Key Concepts:#
Use
os.system()to call.tclor.pyinput files from PythonOutputs go to the path defined in your script
%run(notebook magic) is more limited — it doesn’t accept arguments — soos.system()offers greater flexibility
cwd = os.getcwd()
print('current directory:',cwd)
current directory: /home/jupyter/MyData/_ToCommunityData/OpenSees/TrainingMaterial/training-Computational-Workflows-on-DesignSafe/books/OpenSees-on-DesignSafe/Jupyter_Notebooks_OpenSees
Define Location of Input Files.#
OpsScriptsPath = os.path.expanduser('~/CommunityData/Training/Computational-Workflows-on-DesignSafe/Examples/OpenSees_Basic')
print('OpsScriptsPath:',OpsScriptsPath)
OpsScriptsPath: /home/jupyter/CommunityData/Training/Computational-Workflows-on-DesignSafe/Examples/OpenSees_Basic
Move current working path to user’s home directory#
This way you can save files to your home path – you can’t write to CommunityData
os.chdir(os.path.expanduser('~')) # ~ the tilda is a shortcut to the uers's home path.
cwd = os.getcwd()
print('current directory:',cwd)
current directory: /home/jupyter
Create a temporary directory for our data and move to it#
We want the directory to be within MyData.
tmpDir = 'MyData/tmp_training'
os.makedirs(tmpDir, exist_ok=True)
os.chdir(tmpDir)
cwd = os.getcwd()
print('new current directory:',cwd)
new current directory: /home/jupyter/MyData/tmp_training
# !pip install OpenSeesPy (if needed)
Sequential#
TCL#
OpenSees.exe sequential#
inputFile = 'Ex1a.Canti2D.Push.tcl'
os.system(f'OpenSees {OpsScriptsPath}/{inputFile}')
OpenSees -- Open System For Earthquake Engineering Simulation
Pacific Earthquake Engineering Research Center
Version 3.7.1 64-Bit
(c) Copyright 1999-2016 The Regents of the University of California
All Rights Reserved
(Copyright and Disclaimer @ http://www.berkeley.edu/OpenSees/copyright.html)
Analysis-0 execution done
Analysis-1 execution done
Analysis-2 execution done
Analysis-3 execution done
Analysis-4 execution done
Analysis-5 execution done
Analysis-6 execution done
Analysis-7 execution done
ALL DONE!!!
0
Python#
OpenSeesPy sequential#
inputFile = 'Ex1a.Canti2D.Push.py'
os.system(f'python {OpsScriptsPath}/{inputFile}')
['/home/jupyter/CommunityData/Training/Computational-Workflows-on-DesignSafe/Examples/OpenSees_Basic/Ex1a.Canti2D.Push.py']
Analysis-0 execution done
Analysis-1 execution done
Analysis-2 execution done
Analysis-3 execution done
Analysis-4 execution done
Analysis-5 execution done
Analysis-6 execution done
Analysis-7 execution done
ALL DONE!!!
Process 0 Terminating
0
Parallel#
np=4 # number of processors
TCL#
OpenSeesMP parallel using mpiexec#
inputFile = 'Ex1a.Canti2D.Push.mp.tcl'
os.system(f'mpiexec -np {np} OpenSeesMP {OpsScriptsPath}/{inputFile}')
OpenSees -- Open System For Earthquake Engineering Simulation
Pacific Earthquake Engineering Research Center
Version 3.7.1 64-Bit
(c) Copyright 1999-2016 The Regents of the University of California
All Rights Reserved
(Copyright and Disclaimer @ http://www.berkeley.edu/OpenSees/copyright.html)
pid 2 of np=4 started
pid 3 of np=4 started
pid 1 of np=4 started
pid 0 of np=4 started
pid 2 of 4 Analysis-2 execution done
pid 0 of 4 Analysis-0 execution done
pid 3 of 4 Analysis-3 execution done
pid 1 of 4 Analysis-1 execution done
pid 2 of 4 Analysis-6 execution done
pid 2 ALL DONE!!!
Process Terminating 2
pid 3 of 4 Analysis-7 execution done
pid 3 ALL DONE!!!
Process Terminating 3
pid 0 of 4 Analysis-4 execution done
pid 0 ALL DONE!!!
Process Terminating 0
pid 1 of 4 Analysis-5 execution done
pid 1 ALL DONE!!!
Process Terminating 1
0
f'mpiexec -np {np} OpenSeesMP {OpsScriptsPath}/{inputFile}'
'mpiexec -np 4 OpenSeesMP /home/jupyter/CommunityData/Training/Computational-Workflows-on-DesignSafe/Examples/OpenSees_Basic/Ex1a.Canti2D.Push.mp.tcl'
Python#
OpenSeesPy parallel using mpiexec#
inputFile = 'Ex1a.Canti2D.Push.mpi.py'
os.system(f'mpiexec -np {np} python {OpsScriptsPath}/{inputFile}')
print('If you only see "pid=0" and "np=1", then the MPI implementation failed!')
pid 0 of 1 started
pid 0 of 1 started
pid 0 of 1 started
pid 0 of np=1 Analysis-0 execution done
pid 0 of 1 started
pid 0 of np=1 Analysis-0 execution done
pid 0 of np=1 Analysis-0 execution done
pid 0 of np=1 Analysis-1 execution done
pid 0 of np=1 Analysis-1 execution done
pid 0 of np=1 Analysis-0 execution done
pid 0 of np=1 Analysis-1 execution done
pid 0 of np=1 Analysis-2 execution done
pid 0 of np=1 Analysis-3 execution done
pid 0 of np=1 Analysis-1 execution done
pid 0 of np=1 Analysis-2 execution done
pid 0 of np=1 Analysis-2 execution done
pid 0 of np=1 Analysis-4 execution done
pid 0 of np=1 Analysis-3 execution done
pid 0 of np=1 Analysis-3 execution done
pid 0 of np=1 Analysis-2 execution done
pid 0 of np=1 Analysis-4 execution done
pid 0 of np=1 Analysis-4 execution done
pid 0 of np=1 Analysis-5 execution done
pid 0 of np=1 Analysis-3 execution done
pid 0 of np=1 Analysis-5 execution done
pid 0 of np=1 Analysis-6 execution done
pid 0 of np=1 Analysis-4 execution done
pid 0 of np=1 Analysis-5 execution done
pid 0 of np=1 Analysis-6 execution done
pid 0 of np=1 Analysis-7 execution done
pid 0 of np=1 ALL DONE!!!
pid 0 of np=1 Analysis-5 execution done
pid 0 of np=1 Analysis-6 execution done
pid 0 of np=1 Analysis-7 execution done
pid 0 of np=1 ALL DONE!!!
pid 0 of np=1 Analysis-6 execution done
pid 0 of np=1 Analysis-7 execution done
pid 0 of np=1 ALL DONE!!!
pid 0 of np=1 Analysis-7 execution done
pid 0 of np=1 ALL DONE!!!
If you only see "pid=0" and "np=1", then the MPI implementation failed!
Process 0 Terminating
Process 0 Terminating
Process 0 Terminating
Process 0 Terminating
OpenSeesPy parallel using mpi4py#
inputFile = 'Ex1a.Canti2D.Push.mpi4py.py'
os.system(f'mpiexec -np {np} python {OpsScriptsPath}/{inputFile}')
mpi4py -- python pid 0 of 4 started
mpi4py -- python pid 1 of 4 started
mpi4py -- python pid 3 of 4 started
mpi4py -- python pid 2 of 4 started
pid 2 of np=4 Analysis-2 execution done
pid 1 of np=4 Analysis-1 execution done
pid 0 of np=4 Analysis-0 execution done
pid 3 of np=4 Analysis-3 execution done
pid 1 of np=4 Analysis-5 execution done
pid 1 of np=4 ALL DONE!!!
pid 2 of np=4 Analysis-6 execution done
pid 2 of np=4 ALL DONE!!!
pid 0 of np=4 Analysis-4 execution done
pid 0 of np=4 ALL DONE!!!
pid 3 of np=4 Analysis-7 execution done
pid 3 of np=4 ALL DONE!!!
Process 0 Terminating
Process 0 Terminating
Process 0 Terminating
Process 0 Terminating
0
Run OpenSeesPy script within this notebook#
running the script in this manner maintains all variable definitions and OpenSees-Model Objects (unless wipe was used)
inputFile = 'Ex1a.Canti2D.Push.py'
exec(open(f'{OpsScriptsPath}/{inputFile}').read())
['/opt/conda/lib/python3.12/site-packages/ipykernel_launcher.py', '-f', '/home/jupyter/.local/share/jupyter/runtime/kernel-944c04fb-8fa6-4e2e-8fe4-73f8ef42170b.json']
Command-Line Arguments (argv): ['/opt/conda/lib/python3.12/site-packages/ipykernel_launcher.py', '-f', '/home/jupyter/.local/share/jupyter/runtime/kernel-944c04fb-8fa6-4e2e-8fe4-73f8ef42170b.json']
Analysis-0 execution done
Analysis-1 execution done
Analysis-2 execution done
Analysis-3 execution done
Analysis-4 execution done
Analysis-5 execution done
Analysis-6 execution done
Analysis-7 execution done
ALL DONE!!!
Variables & OpenSees model have been preserved:
# OpenSees Model have been preserved (unless wipe has been used)
print('OpenSees Model')
ops.printModel()
OpenSees Model
Current Domain Information
Current Time: 6.19189
Committed Time: 6.19189
NODE DATA: NumNodes: 2
numComponents: 2
Node: 1
Coordinates : 0 0
Disps: 0 0 0
unbalanced Load: 0 0 0
reaction: -12383.8 2000 5.94422e+06
ID : -1 -1 -1
Node: 2
Coordinates : 0 480
Disps: 100 -6.30865e-08 -0.3125
unbalanced Load: 12383.8 -2000 0
reaction: -1.81899e-12 2.27374e-13 -4.65661e-10
Mass :
5.18 0 0
0 0 0
0 0 0
Rayleigh Factor: alphaM: 0
Rayleigh Forces: 0 0 0
ID : 0 1 2
ELEMENT DATA: NumEle: 1
numComponents: 1
ElasticBeam2d: 1
Connected Nodes: 1 2
CoordTransf: 1
mass density: 0, cMass: 0
release code: 0
End 1 Forces (P V M): 2000 12383.8 5.94422e+06
End 2 Forces (P V M): -2000 -12383.8 -4.65661e-10
SP_Constraints: numConstraints: 3
numComponents: 3
SP_Constraint: 0 Node: 1 DOF: 1 ref value: 0 current value: 0 initial value: 0
SP_Constraint: 1 Node: 1 DOF: 2 ref value: 0 current value: 0 initial value: 0
SP_Constraint: 2 Node: 1 DOF: 3 ref value: 0 current value: 0 initial value: 0
Pressure_Constraints: numConstraints: 0
numComponents: 0
MP_Constraints: numConstraints: 0
numComponents: 0
LOAD PATTERNS: numPatterns: 2
numComponents: 2
Load Pattern: 1
Scale Factor: 1
Linear Series: constant factor: 1
Nodal Loads:
numComponents: 1
Nodal Load: 2 load : 0 -2000 0
Elemental Loads:
numComponents: 0
Single Point Constraints:
numComponents: 0
Load Pattern: 2
Scale Factor: 1
Linear Series: constant factor: 1
Nodal Loads:
numComponents: 1
Nodal Load: 2 load : 2000 0 0
Elemental Loads:
numComponents: 0
Single Point Constraints:
numComponents: 0
PARAMETERS: numParameters: 0
numComponents: 0
Plot analysis results#
for any of the above analyses
#pick any case
dataDir = 'DataTCL'
Lcol = 400
# check paths:
print('relative path:',os.path.expanduser(dataDir))
print('absolute path:',os.path.abspath(dataDir))
print('current directory:',os.getcwd())
relative path: DataTCL
absolute path: /home/jupyter/MyData/tmp_training/DataTCL
current directory: /home/jupyter/MyData/tmp_training
plt.close('all')
fname3 = f'{dataDir}/DFree_Lcol{Lcol}.out'
dataDFree = numpy.loadtxt(fname3)
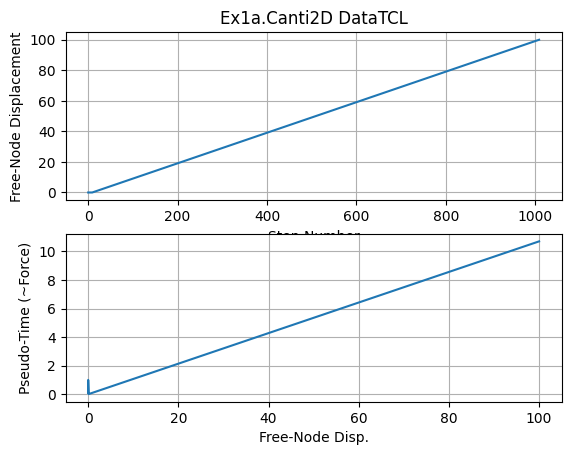
plt.subplot(211)
plt.title(f'Ex1a.Canti2D {dataDir}')
plt.grid(True)
plt.plot(dataDFree[:,1])
plt.xlabel('Step Number')
plt.ylabel('Free-Node Displacement')
plt.subplot(212)
plt.grid(True)
plt.plot(dataDFree[:,1],dataDFree[:,0])
plt.xlabel('Free-Node Disp.')
plt.ylabel('Pseudo-Time (~Force)')
plt.savefig(f'{dataDir}/Response.jpg')
plt.show()
print(f'plot saved to {dataDir}/Response_Lcol{Lcol}.jpg')
print('End of Run: Ex1a.Canti2D.Push.py.ipynb')
plot saved to DataTCL/Response_Lcol400.jpg
End of Run: Ex1a.Canti2D.Push.py.ipynb
print('done!')
done!
